Probability and Statistics: A Geometric Foundation
This lecture includes interactive SymPy cells that verify key results symbolically. SymPy is a Python library for symbolic mathematics. Loading it fetches a WebAssembly Python runtime (approx. 15 MB, cached after first load). You can also load it on demand from any code cell below.
This article includes interactive NumPy cells that verify the numerical examples in each section. NumPy runs in a WebAssembly Python runtime (approx. 5 MB, cached after first load). You can also load it on demand from any code cell below.
1. Probability as measure
Most first encounters with probability present it as a set of rules for computing with fractions: count the favourable outcomes, divide by the total, apply Bayes' formula. This machinery works well enough for coins and dice, but it quietly falls apart the moment we ask about continuous quantities, infinite sample spaces, or the convergence of sequences of random variables. The reason is that probability is, at its core, a theory of measurement. The question "what is the probability of an event?" is really the question "what is the size of a set?", and answering it properly requires the same infrastructure that underpins the Lebesgue integral: -algebras, measures, and measurable functions.
This section rebuilds the foundations from that starting point. The payoff is not abstraction for its own sake. Once probability is grounded in measure theory, results like the law of large numbers and the central limit theorem become theorems about integrals, and the passage from discrete to continuous distributions requires no new definitions, only a change of reference measure. The Radon-Nikodym derivative, which we introduce at the end of this section, makes that change of reference precise and opens the door to the geometric perspective that runs through the rest of the post.
We begin with the two structures that any probabilistic model requires: a way to specify which events are observable, and a way to assign sizes to them.
Let be a non-empty set. A collection of subsets of is a -algebra if it satisfies:
- .
- If , then .
- If , then .
The pair is called a measurable space.
Condition (iii) is the critical one: closure under countable unions, not merely finite ones. This is what separates -algebras from the simpler algebras of sets and is precisely the condition needed to take limits. Without it, we could not define convergence of events, and probability theory would be unable to make asymptotic statements.
Every set admits at least two sigma-algebras: the trivial sigma-algebra , in which only the certain and impossible events are observable, and the power set , in which every subset is observable. For uncountable spaces like , the power set is too large to support a useful measure, and we work instead with the Borel sigma-algebra , generated by the open intervals. The choice of sigma-algebra encodes what we can observe: a coarser sigma-algebra means less information.
Let be a measurable space. A function is a probability measure if:
- .
- Countable additivity: for any sequence of pairwise disjoint sets in ,
The triple is called a probability space.
The axioms are Kolmogorov's (1933), and their power lies in what they leave unspecified. They say nothing about equally likely outcomes, nothing about frequencies, nothing about subjective beliefs. They say only that probability is a normalised, countably additive set function. Every interpretation of probability (frequentist, Bayesian, decision-theoretic) is a commitment about what means; the measure-theoretic framework is agnostic.
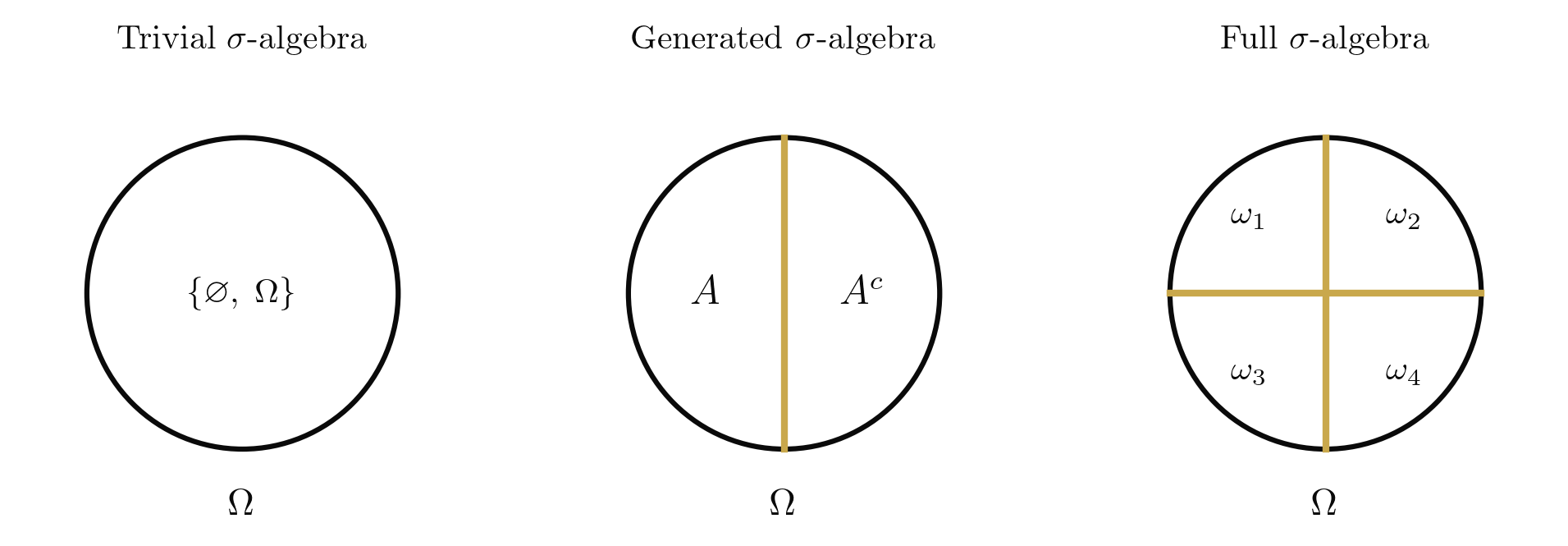
With the probability space in hand, we can formalise what it means to assign a numerical value to each outcome.
Let be a measurable space. A random variable is a measurable function , where is the Borel sigma-algebra on . That is, for every Borel set ,
The measurability condition [eq:measurability] ensures that asking "what is the probability that falls in ?" is a well-posed question: the preimage belongs to , so can evaluate it. A function that is not measurable can produce sets whose probability is undefined, which is precisely the pathology that arises when one tries to assign probabilities to all subsets of (the Vitali construction).
Let be a random variable on the probability space . The expectation of is defined as the Lebesgue integral of with respect to :
constructed in three stages. First, for a simple function with , define . Second, for a non-negative measurable , define the integral as the supremum over all simple functions bounded above by . Third, for a general , decompose where and , and set , provided at least one of these is finite.
The three-stage construction mirrors the Lebesgue integral exactly, because that is what expectation is. The only difference from an analyst's perspective is that the underlying measure is a probability measure rather than the Lebesgue measure. This identification is the central simplification of the measure-theoretic approach: every theorem about Lebesgue integrals (dominated convergence, monotone convergence, Fubini-Tonelli) applies immediately to expectations.
The following theorem illustrates why countable additivity, rather than mere finite additivity, is the right axiom. It lets us compute probabilities of limits from limits of probabilities.
Let be a probability space. If is an increasing sequence of events with , then
Symmetrically, if is a decreasing sequence with , then .
Proof
Define and for . The sets are pairwise disjoint, and . By countable additivity [eq:countable-additivity],
where the last equality holds because by finite additivity (a consequence of countable additivity) and the fact that .
The decreasing case follows by applying the increasing case to the complements and using .
Let us see all of these definitions at work in the simplest non-trivial probability space.
Take and (the power set). For a parameter , define by
This determines on all of by additivity: and . The resulting probability space is the Bernoulli space .
The identity function defined by is trivially measurable (every subset of is in ). Its expectation is
and its variance is
The Bernoulli space is minimal, but it already contains the essential structure: a sample space, a -algebra encoding what we can observe, a measure encoding our beliefs, and a random variable converting outcomes to numbers. Every probability model, no matter how complex, is built from the same ingredients.
1.1. Changing the reference: the Radon-Nikodym derivative
So far we have worked with a single probability measure . In practice we constantly compare two measures on the same measurable space: a prior and a posterior, a model and the data-generating distribution, a risk-neutral and a physical measure. The Radon-Nikodym derivative makes such comparisons precise.
Let and be -finite measures on . We say is absolutely continuous with respect to , written , if for every ,
When , the Radon-Nikodym theorem ([radon-nikodym-thm]) guarantees the existence of a measurable function such that for all ,
The function , unique -almost everywhere, is called the Radon-Nikodym derivative of with respect to and is written .
Let and be -finite measures on with . Then there exists a measurable function such that
Moreover, is unique -almost everywhere.
Proof
We sketch the proof via the Riesz representation theorem, following the approach of von Neumann.
Define , so that both and are absolutely continuous with respect to . For any , the map
defines a bounded linear functional on (bounded because ). By the Riesz representation theorem, there exists such that
Taking for gives . Since for all , we have -almost everywhere. Similarly, implies -a.e.
On the set where , we can write , which rearranges to , giving . Setting on and verifying that (which follows from ) yields the desired density. Uniqueness follows from the fact that if two functions agree -a.e., their integrals over every measurable set coincide.
The Radon-Nikodym derivative is the infinitesimal ratio of -volume to -volume. When is the Lebesgue measure on , the derivative is the familiar probability density function from a first course in statistics. But the Radon-Nikodym perspective reveals something that the elementary treatment obscures: a density is not an intrinsic property of a distribution. It is a relative quantity, depending on the choice of reference measure. Change the reference, and the density changes with it. This relativity is the first hint that probability has a geometric character, one that will become explicit when we study the Fisher metric in later sections.
1.2. The geometric thread: families as curves
Consider the family of all Bernoulli distributions parametrised by . Each value of defines a probability measure on , and these measures live in the probability simplex
The Bernoulli family traces a curve in this simplex: . The simplex is the ambient space of all distributions on a finite sample space, and parametric families are curves or surfaces within it. This geometric perspective, viewing families of distributions as geometric objects, will deepen throughout this post. For now, note that the curve passes through every point in , so the Bernoulli family is a full parametrisation of the simplex. Richer sample spaces will give higher-dimensional simplices, and the parametric families we study will typically be lower-dimensional submanifolds.
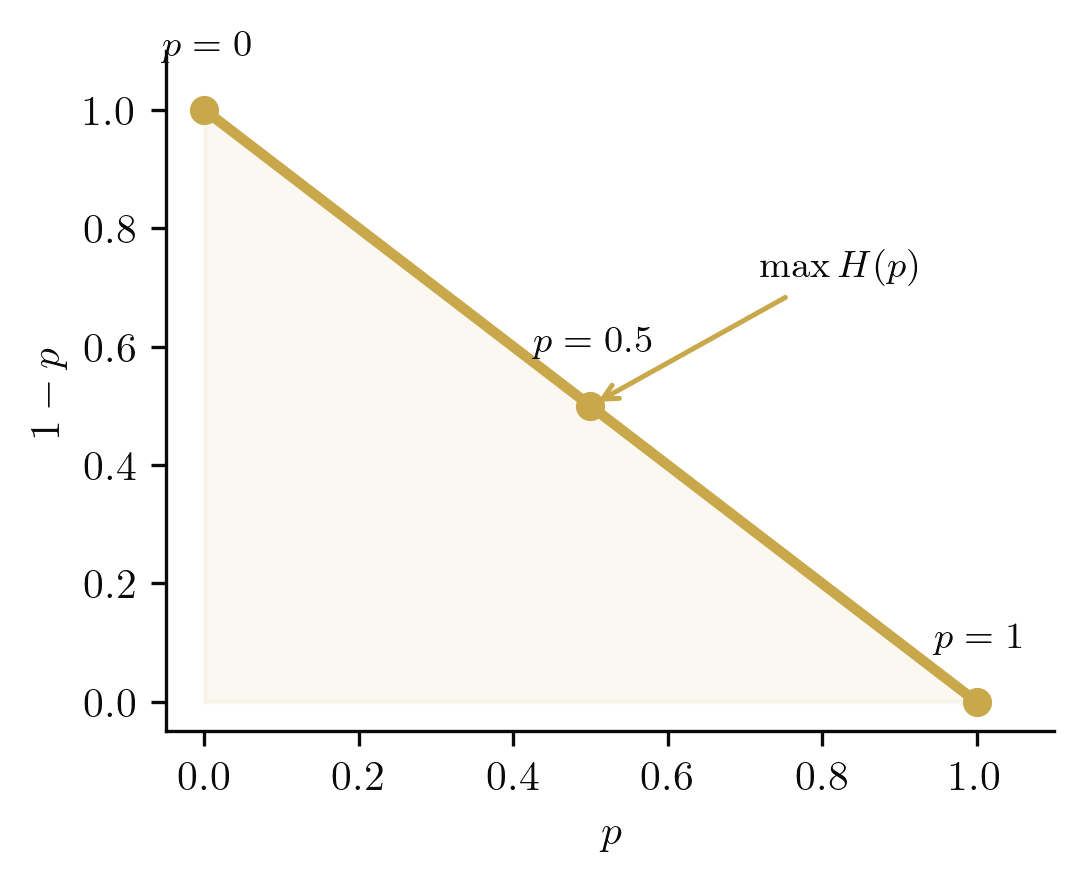
A natural question arises: which point on this curve is "most uncertain"? The answer leads to entropy.
The Shannon entropy of is
This function is maximised at , where bit, and vanishes at and (certainty). The entropy quantifies our uncertainty about the outcome: high entropy means the result is unpredictable, while zero entropy means the outcome is determined before we observe it. In the simplex picture, the maximum-entropy point sits at the midpoint of , equidistant from both vertices.
from sympy import symbols, log, simplify, Rational
p = symbols('p', positive=True)
# Expectation of Ber(p)
EX = 0*(1-p) + 1*p
print(f"E[X] = {EX}")
# Variance
VarX = simplify(EX*(1-EX)) # E[X^2] - (E[X])^2 = p - p^2
print(f"Var(X) = {VarX}")
# Shannon entropy (base 2)
H = simplify(-p*log(p, 2) - (1-p)*log(1-p, 2))
print(f"H(p) = {H}")
# Value at p = 1/2
print(f"H(1/2) = {H.subs(p, Rational(1,2))} bits")Bernoulli expectation, variance, and entropy
2. Estimation theory
With the measure-theoretic foundations in place, we turn to the central question of statistics: given observed data, what can we infer about the mechanism that generated it? This requires a formal notion of a statistical model and a systematic theory of estimation.
2.1. Statistical models and likelihood
The starting point is to specify what we mean by a "family of possible data-generating processes." A statistical model formalises this idea by indexing probability measures with a parameter.
A statistical model is a family of probability measures on a measurable space , indexed by a parameter space . If each has a density with respect to a common dominating measure , the model is called a parametric model.
The parametric assumption, that the family is smoothly indexed by a finite-dimensional parameter, is what makes classical estimation theory tractable. The dominating measure is typically Lebesgue measure (for continuous models) or counting measure (for discrete models). Its role is to allow us to work with densities rather than abstract measures.
Given a parametric model, an observation singles out a function on the parameter space: how "compatible" is each with what we saw?
Given observed data , the likelihood function is
the density evaluated at , viewed as a function of . The log-likelihood is
For i.i.d. observations , the log-likelihood is
The likelihood is emphatically not a probability distribution over : it does not integrate to one, and it carries no probabilistic interpretation on the parameter space (that would require a prior, which we defer to the Bayesian section). It is simply a score of compatibility. The log-likelihood is preferred in practice because products become sums, which are numerically stable and analytically convenient.
The most natural estimation principle is to choose the parameter value that makes the observed data most likely.
The maximum likelihood estimator (MLE) is
When the log-likelihood is differentiable, the MLE solves the score equation:
The notation deserves a moment of attention, since it appears throughout estimation theory. Consider a real-valued function defined on some set .
The maximum of is the largest value attained:
The argmax of is the point (or set of points) at which that maximum is attained:
In other words, returns how large the function gets; returns where it gets there. When the maximiser is unique (as it typically is for log-likelihoods of regular models), we write as though it returns a single element rather than a set.
The MLE is a function of the data, hence a random variable. Its properties (consistency, efficiency, asymptotic normality) are among the deepest results in estimation theory, and we will establish them below after developing the necessary machinery.
2.2. Sufficient statistics
Not all functions of the data are equally useful. Some compress the data without losing any information about the parameter; others discard relevant information. The notion of sufficiency makes this precise.
A statistic is sufficient for if the conditional distribution of given does not depend on . Intuitively, captures all the information in the data about : once is known, the remaining randomness in is pure noise.
Verifying sufficiency directly from the conditional distribution is often impractical. The following factorisation criterion provides a clean, checkable condition.
A statistic is sufficient for if and only if the density factors as
for some functions and , where does not depend on .
Proof
If is sufficient, then . The first factor does not depend on (by sufficiency); call it . The second depends on only through ; call it .
If , then
which does not depend on .
2.3. Exponential families
Sufficiency becomes particularly elegant in the context of exponential families, which encompass most of the standard parametric models and admit a rich geometric structure.
A parametric family is an exponential family if the density can be written
where is the natural parameter, is the sufficient statistic, is the log-partition function (ensuring normalisation), and is the base measure.
The factorisation theorem immediately tells us that is sufficient for . The log-partition function is convex (since it is the logarithm of a moment-generating function), and its derivatives encode the moments of the sufficient statistic.
For , write
So (log-odds), , , and .
For , the density is . Rewrite as
where , , , and
The following result shows that the log-partition function is a generating function for the cumulants of the sufficient statistic.
For an exponential family with log-partition function ,
Proof
The normalisation condition is
so . Differentiating both sides with respect to :
Dividing by yields . Differentiating again gives the covariance:
Since is convex, the Hessian is positive semi-definite, consistent with the covariance matrix being non-negative definite.
2.4. Fisher information
The curvature of the log-likelihood at the true parameter value quantifies how much information the data carries about . This is formalised by the Fisher information.
The Fisher information at is
The quantity is the score function. Since (the score has mean zero under the model), the Fisher information equals the variance of the score: . Under regularity conditions (interchange of differentiation and integration),
For a multi-parameter model with , the Fisher information matrix has entries
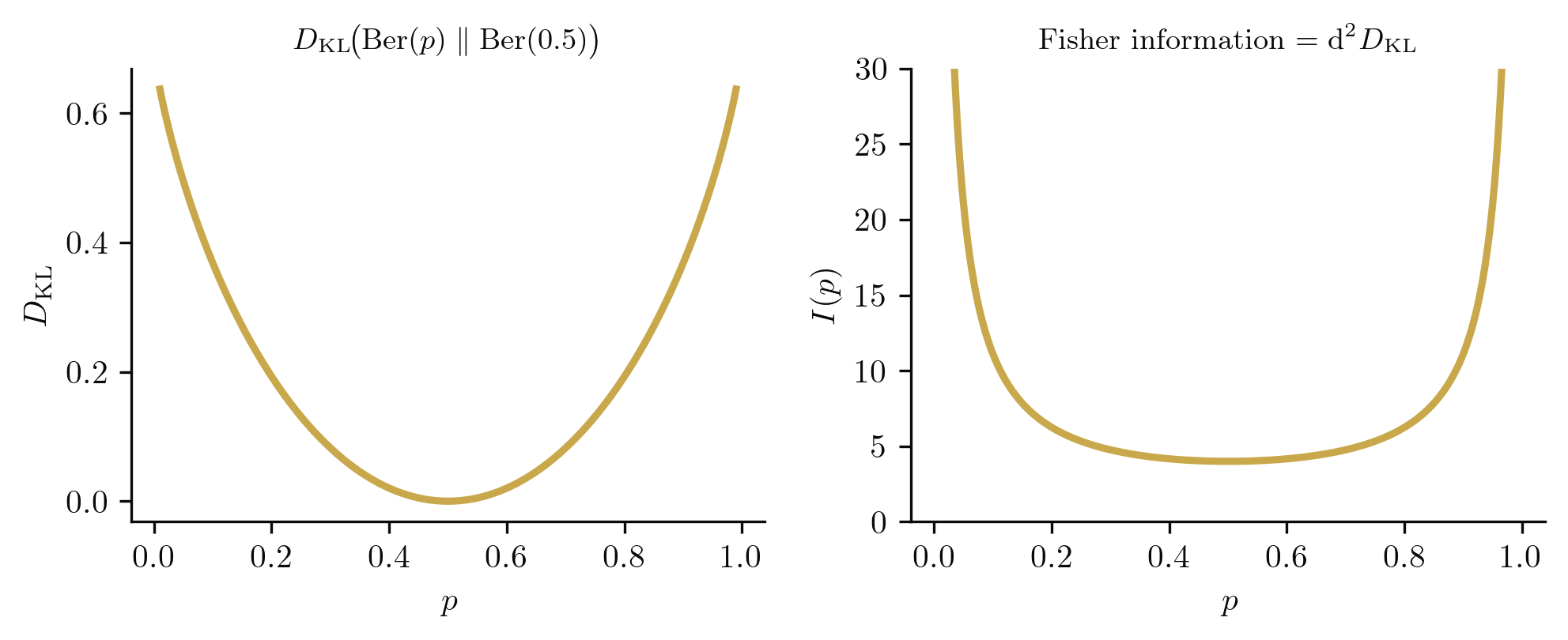
For , the log-density is . The score is
Since , we obtain
Note how diverges as or : near certainty, each observation is maximally informative.
For with , the Fisher information matrix is
The diagonal structure reflects the orthogonality of the location and scale parameters. Reparametrising to yields .
from sympy import symbols, log, diff, simplify, Function
p, x = symbols('p x')
# Log-likelihood for single Bernoulli observation
logp = x*log(p) + (1-x)*log(1-p)
score = diff(logp, p)
print(f"Score: {score}")
# Fisher info: -E[d^2/dp^2 log p] = -E[second derivative]
d2 = diff(logp, p, 2)
# E[x] = p, E[1-x] = 1-p
I_p = simplify(-d2.subs(x, p)) # E[d2] by replacing x with E[x]=p
print(f"I(p) = {I_p}")Fisher information for the Bernoulli family
from sympy import symbols, log, diff, simplify, pi, sqrt, Matrix
x, mu, sigma = symbols('x mu sigma', real=True, positive=True)
# Log-density of N(mu, sigma^2)
logp = -log(sigma*sqrt(2*pi)) - (x - mu)**2 / (2*sigma**2)
# Score components
s_mu = diff(logp, mu)
s_sig = diff(logp, sigma)
print(f"Score (mu): {simplify(s_mu)}")
print(f"Score (sigma): {simplify(s_sig)}")
# Fisher info: I_ij = E[s_i * s_j]
# Use E[x] = mu, E[(x-mu)^2] = sigma^2, E[(x-mu)^4] = 3*sigma^4
from sympy import Rational
I_11 = simplify(s_mu**2)
I_11 = I_11.subs((x-mu)**2, sigma**2) # take expectation
I_22 = simplify(s_sig**2)
I_22 = I_22.subs((x-mu)**4, 3*sigma**4).subs((x-mu)**2, sigma**2)
print(f"I_11 = E[s_mu^2] = {simplify(I_11)}")
print(f"I_22 = E[s_sigma^2] = {simplify(I_22)}")Fisher information matrix for the Normal family
The Fisher information admits a compelling geometric interpretation that will become central in Section 4.
The Fisher information measures the curvature of the log-likelihood at . A sharply peaked log-likelihood (high ) means the data is informative about ; a flat log-likelihood (low ) means the data tells us little. For the Bernoulli family, diverges at the boundary of the parameter space: the curve traced by the Bernoulli family in the simplex bends sharply near and . For the Normal family, the Fisher information matrix encodes a two-dimensional geometry on the plane. In Section 4, we will show that this matrix is a Riemannian metric, and the "curvature" here becomes literal geometric curvature.
2.5. Bias, variance, and the Cramer-Rao bound
Before establishing the optimality of the MLE, we need a framework for comparing estimators. The fundamental decomposition of estimation error separates systematic error (bias) from random fluctuation (variance).
For an estimator of :
- Bias: .
- Mean squared error: .
An estimator is unbiased if for all .
The MSE decomposition reveals a fundamental tension: reducing bias (e.g. by shrinkage) may increase variance, and vice versa. The "best" estimator depends on which trade-off one is willing to accept. Within the class of unbiased estimators, however, there is a sharp lower bound on the variance.
An unbiased estimator is efficient if
i.e. it achieves the Cramer-Rao lower bound with equality.
For any unbiased estimator of based on i.i.d. observations,
Proof
Consider a single observation. The score satisfies and . For any unbiased estimator with , differentiate both sides with respect to :
Since , this gives , i.e. (using ). By the Cauchy-Schwarz inequality:
Therefore . For i.i.d. observations, the total Fisher information is , giving .
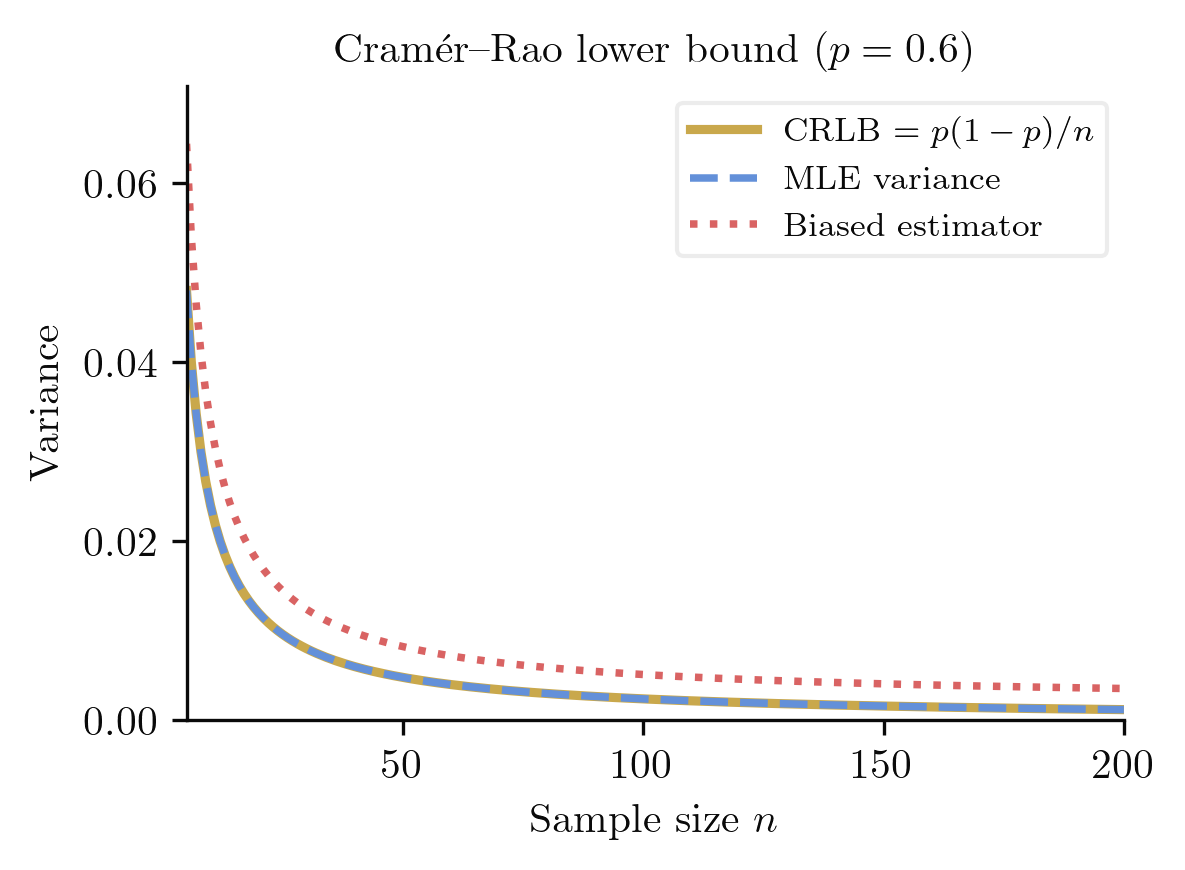
The Cramer-Rao bound is tight in exponential families: for any exponential family, the MLE of achieves the bound exactly. Outside exponential families, the bound may not be attainable by any unbiased estimator, but the MLE still achieves it asymptotically.
2.6. Asymptotic efficiency of the MLE
The deepest result in classical estimation theory is that the MLE is, in a precise sense, the best estimator one can construct from large samples.
Under regularity conditions (the parameter space is open, the support of does not depend on , and ), the MLE is consistent () and asymptotically normal:
In particular, the MLE is asymptotically efficient: its variance achieves the Cramer-Rao lower bound in the limit.
Proof sketch
Taylor-expand the score around the true parameter . At the MLE, the score vanishes:
By the law of large numbers, . By the central limit theorem, . Solving for :
Making this argument rigorous requires uniform convergence of the Taylor remainder, which follows from the regularity conditions.
This theorem closes the circle: the Fisher information sets a fundamental limit on estimation precision (Cramer-Rao), and the MLE saturates that limit for large samples. The connection to geometry is immediate. The asymptotic covariance of the MLE is the inverse of the Fisher information matrix, which we will identify in Section 4 as the inverse of the Riemannian metric on the statistical manifold. Estimation theory and differential geometry are, in this sense, two perspectives on the same structure.
3. Inference and testing
Estimation tells us what to believe about a parameter; testing tells us what we can rule out. Hypothesis testing provides a formal framework for decisions under uncertainty, and its mathematical structure flows directly from the likelihood theory we have developed.
The central question shifts from "what is the best estimate of ?" to "is a particular value of compatible with the data?" To answer this, we need a decision procedure with quantifiable error guarantees.
3.1. The testing framework
A hypothesis test consists of:
- A null hypothesis and an alternative hypothesis , where and partition the parameter space ,
- A test statistic , and
- A critical region such that we reject if .
Any decision procedure that divides the sample space into "reject" and "fail to reject" regions can make two kinds of mistakes. The entire theory of testing is built around controlling these errors.
The size (Type I error rate) of a test is
The Type II error rate is
The power function is
A good test has small (rarely rejects a true null) and large power (reliably detects true alternatives). These goals are in tension: shrinking the critical region reduces but also reduces power. The Neyman-Pearson framework resolves this by fixing and maximising power.
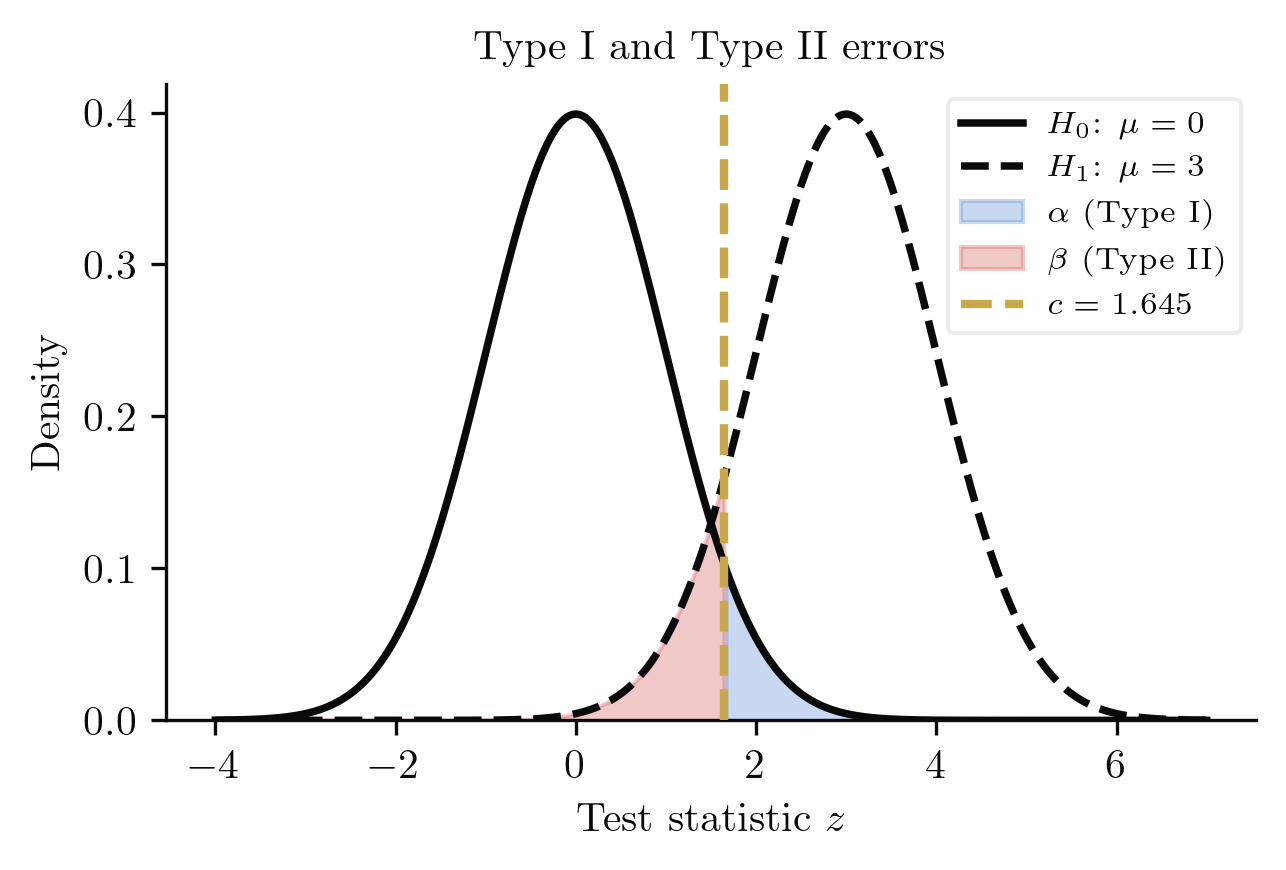
Rather than choosing a significance level in advance, we can summarise the evidence against with a single number.
The p-value is the probability, under , of observing a test statistic at least as extreme as the one realised:
Under , the p-value is uniformly distributed on .
The uniformity statement is worth emphasising: if is true, the p-value is no more likely to fall below 0.05 than below any other threshold of the same width. This is what makes the p-value a valid calibration of evidence.
3.2. Optimal tests
Given the trade-off between Type I and Type II errors, a natural question arises: among all tests of a given size, which has the greatest power? The Neyman-Pearson lemma gives a complete answer for simple hypotheses.
For testing the simple hypothesis against the simple alternative at size , the most powerful test rejects when
where is chosen so that .
Proof
Let denote the Neyman-Pearson test (reject when ) and let be any other test of size . We need to show .
Consider the integral
In the region , we have and , so since .
In the region , we have and , so
since and .
Therefore the integral is non-negative. Expanding:
The right-hand side is since has size . Hence , and the Neyman-Pearson test is most powerful.
The lemma tells us that the likelihood ratio is the sufficient statistic for distinguishing between two simple hypotheses. All the information relevant to the decision is captured in .
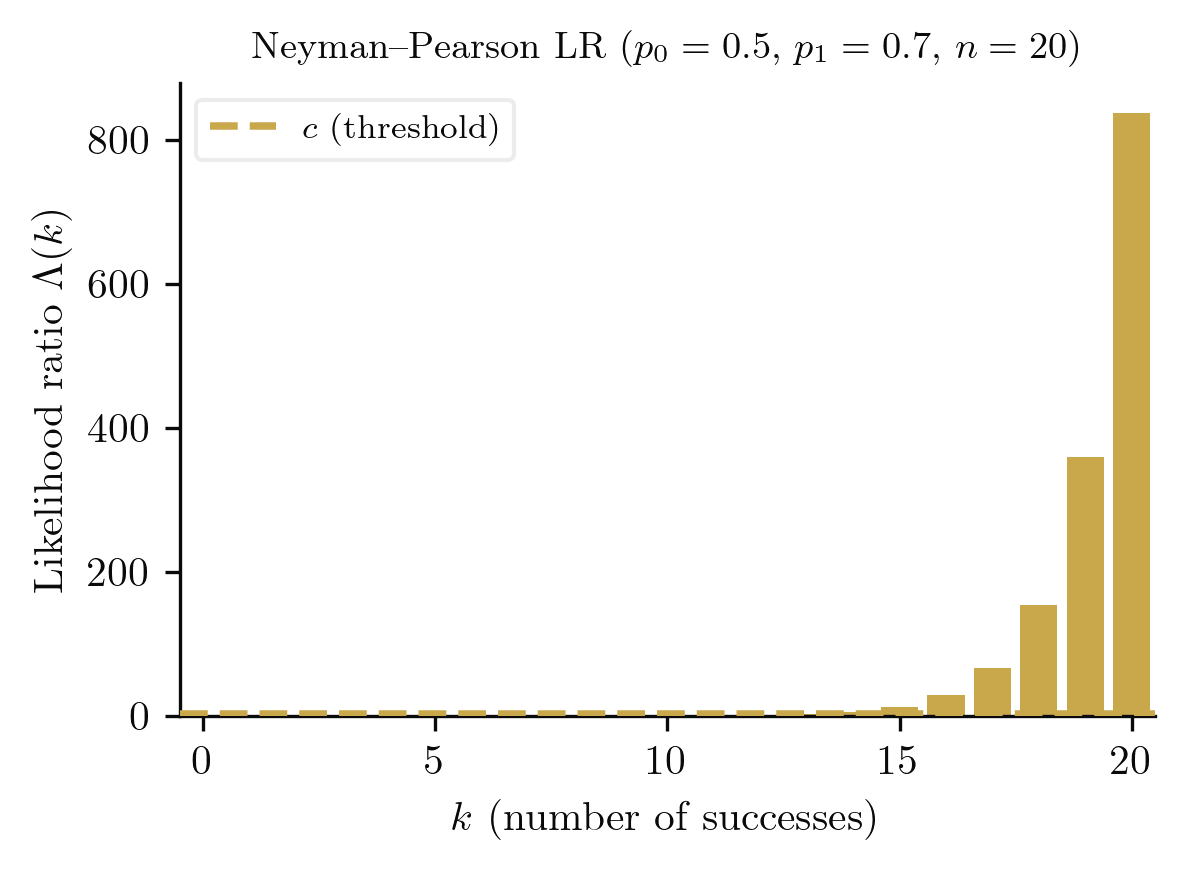
The Neyman-Pearson lemma handles simple vs simple hypotheses. For composite alternatives (e.g., ), we need a stronger condition on the statistical model.
If a family has monotone likelihood ratio in , then the uniformly most powerful (UMP) test of against at size rejects when , where satisfies .
Proof
Fix any . By the Neyman-Pearson lemma, the most powerful test of vs rejects when
for some threshold . Since the family has monotone likelihood ratio in , the ratio is non-decreasing in . Therefore
for some that depends only on (through ) and not on . Since the rejection region does not depend on the specific alternative , the test is UMP against the entire composite alternative .
Exponential families have monotone likelihood ratio in their natural sufficient statistics, so one-sided UMP tests exist for all standard exponential family models.
3.3. Large-sample testing
When exact UMP tests are unavailable (as with two-sided or multi-parameter problems), the likelihood ratio test provides an asymptotically optimal alternative.
For testing , where is an -dimensional subspace of the -dimensional parameter space , the likelihood ratio statistic
converges in distribution to under as .
Proof sketch
Taylor-expand the log-likelihood around the unrestricted MLE :
where the first-order term vanishes because . Therefore
By the asymptotic normality of the MLE, . The quadratic form , where , equals for . Under the constraint that coordinates are free, this quadratic form follows a distribution.
Wilks' theorem is remarkably practical: it provides an asymptotic null distribution that does not depend on the true parameter value, only on the dimensions of the full and restricted parameter spaces. This universality is why the likelihood ratio test is the default in most applied settings.
3.4. Confidence regions
Testing and estimation are two sides of the same coin. The connection runs through confidence regions.
A confidence region for is a random set such that
The construction of confidence regions often relies on quantities whose distributions are parameter-free.
A pivotal quantity is a function of the data and the parameter whose distribution does not depend on . If has known distribution , then
is a confidence region.
The fundamental connection between testing and confidence regions is captured by the following duality.
There is a duality between hypothesis tests and confidence regions:
Proof
By construction, if and only if we fail to reject at level . When is the true parameter, the test rejects with probability at most . Therefore
This holds for every , so is a valid confidence region.
This duality means that every test at level generates a confidence region (by inverting the acceptance region), and every confidence region generates a family of tests (by checking membership). The two frameworks carry exactly the same information.
3.5. Examples
Consider testing against with coin flips. The exact binomial test rejects if or , where and are chosen so the total rejection probability under is at most .
The likelihood ratio approach rejects when
For moderate , the two tests give similar (but not identical) rejection regions. The exact test controls size precisely; the LRT relies on the approximation from Wilks' theorem.
With known variance , the z-test uses the pivotal quantity
and rejects when .
With unknown variance, we replace with the sample standard deviation , yielding
That follows a -distribution (rather than something more complicated) rests on a structural fact about the Normal: and are independent, a consequence of the sufficiency of and the orthogonal decomposition of the sample space. The confidence interval is precisely the inversion of this test, illustrating the confidence-test duality.
For jointly, the confidence ellipse is centred at and shaped by the observed Fisher information. In the standard Euclidean metric on -space, this ellipse treats a step of size in the same whether or .
But the Fisher information matrix tells a different story. Recall that for the Normal family, the Fisher metric is
which says that -space is not flat. At small (where is large), a step of fixed coordinate size traverses far more statistical distance than the same step at large . The "correct" confidence regions should account for this curvature.
The confidence ellipses above are computed using the Euclidean distance in space. But the Fisher information matrix says this space is not flat. In the Euclidean metric, a step of size in is the same whether or . In the Fisher metric, a step at small (where is large) covers far more statistical distance.
The toggle above reveals the difference: Fisher-metric confidence regions are elongated where the parameter space is "flat" (large ) and compressed where it is "curved" (small ). We will make this precise in the next section.
3.6. Computational verification
The coverage probability of a confidence interval is a frequentist guarantee: over repeated samples, the interval should contain the true parameter at least of the time. We can verify this directly.
import numpy as np
np.random.seed(42)
p_true, n, alpha = 0.6, 30, 0.05
z = 1.96
n_sim = 10000
covers = 0
for _ in range(n_sim):
x = np.random.binomial(n, p_true)
p_hat = x / n
se = np.sqrt(p_hat * (1 - p_hat) / n)
lo, hi = p_hat - z * se, p_hat + z * se
if lo <= p_true <= hi:
covers += 1
print(f"Wald CI coverage: {covers/n_sim:.4f} (nominal: {1-alpha:.2f})")
print(f"(n={n}, p={p_true}, {n_sim} simulations)")Monte Carlo verification of Wald interval coverage
The Wald interval is known to undercover for moderate and extreme , a phenomenon that motivates alternatives like the Wilson interval. The gap between nominal and actual coverage is itself a statistical quantity that we can study.
The power curve of a test visualises how detection probability varies with the true parameter value. At , the power equals the size ; away from , it should increase toward 1.
import numpy as np
from scipy import stats
n = 20
alpha = 0.05
# Two-sided exact binomial test of H0: p = 0.5
p_values = np.linspace(0.01, 0.99, 50)
power = np.zeros_like(p_values)
for i, p in enumerate(p_values):
# Power = P(reject H0 | true p)
pw = 0
for k in range(n + 1):
pval = stats.binom_test(k, n, 0.5, alternative='two-sided')
if pval <= alpha:
pw += stats.binom.pmf(k, n, p)
power[i] = pw
print("p power")
for p, pw in zip(p_values[::5], power[::5]):
print(f"{p:.2f} {pw:.4f}")
print(f"\nPower at p=0.5: {power[24]:.4f} (should be ~alpha={alpha})")
print(f"Power at p=0.8: {power[39]:.4f}")Power curve for the Bernoulli two-sided test
The power curve is symmetric about (since the test is two-sided) and increases monotonically as moves away from the null. The rate of increase depends on : larger samples yield steeper power curves, concentrating the "uncertain" region into a narrower band around .
4. The geometric synthesis
Throughout this post, we have spoken informally of the "curvature" of the log-likelihood, the "geometry" of the parameter space, and the "shape" of confidence regions. It is time to make these metaphors precise. The parameter space of a statistical model carries a natural Riemannian structure, the Fisher metric, that encodes the intrinsic geometry of statistical inference. In this section, we construct this geometry from first principles.
4.1. The manifold structure
A smooth manifold of dimension is a second-countable Hausdorff topological space together with a collection of pairs (called charts), where each is an open subset of and is a homeomorphism onto an open subset of , such that:
- The charts cover : .
- For any two charts with , the transition map is smooth ().
The parameter space of a regular statistical model is a smooth manifold: openness in provides a single global chart, but the geometric viewpoint allows for parameter spaces with nontrivial topology (circles, spheres, positive-definite cones).
A tangent vector at is an equivalence class of smooth curves with , where two curves are equivalent if
in any chart containing . The tangent space is the set of all tangent vectors at ; it is a real vector space of dimension . In local coordinates , every tangent vector can be written
where the Einstein summation convention is in force: repeated upper and lower indices are summed from to .
4.2. The Riemannian structure
A manifold on its own has no notion of distance or angle. That structure is imposed by a metric.
A Riemannian metric on is a smooth assignment of a positive-definite inner product on each tangent space . In local coordinates,
where are smooth functions and the matrix is symmetric and positive-definite at every . The pair is called a Riemannian manifold.
Where does such a metric come from in statistics? The answer is the Fisher information.
Let be a regular statistical model. The Fisher information matrix defines a Riemannian metric on the statistical manifold by
This is a -tensor field on , called the Fisher-Rao metric or simply the Fisher metric. The arc length element is
We must verify that this definition actually yields a Riemannian metric: the key property is positive-definiteness.
On a regular statistical model, is positive-definite at every .
Proof
For any tangent vector , we compute
This is the expectation of a squared random variable, so . Equality holds if and only if almost surely under . For a regular model, the score functions are linearly independent as random variables (this is part of the regularity conditions guaranteeing that the Fisher matrix is nonsingular). Therefore implies .
The next theorem is the reason the Fisher metric deserves to be called geometric: it does not depend on the coordinate system.
Under a smooth reparametrisation , the Fisher matrix transforms as
This is the transformation law of a -tensor, confirming that the Fisher metric is a geometric object independent of the coordinate system.
Proof
By the chain rule applied to the score function,
Substituting into the definition of the Fisher metric in the new coordinates:
where the Jacobian factors exit the expectation because they are deterministic functions of the parameters.
4.3. Connection, geodesics, and curvature
A Riemannian metric determines a canonical way to differentiate vector fields along curves: the Levi-Civita connection. Its components in local coordinates are the Christoffel symbols.
The Christoffel symbols of the Levi-Civita connection are
where denotes the entry of the inverse matrix of .
A geodesic is a curve that parallel-transports its own tangent vector. In local coordinates, it satisfies the system of second-order ODEs
Geodesics are the curves of zero acceleration; they generalise straight lines to curved spaces. The geodesic distance between two points is the infimum of the arc lengths of all smooth curves connecting them.
The Riemann curvature tensor is defined in local coordinates by
For a two-dimensional manifold, the Gaussian curvature is the single scalar
The sign of determines the local geometry: (spherical), (flat), (hyperbolic).
4.4. The Fisher metric as infinitesimal KL divergence
The deepest justification for the Fisher metric is not algebraic but information-theoretic: it arises as the leading-order term in the Taylor expansion of the Kullback-Leibler divergence.
The Kullback-Leibler divergence (or relative entropy) between distributions and is
It is non-negative ( iff ) but not symmetric and does not satisfy the triangle inequality, so it is not a metric in the topological sense.
For an infinitesimal displacement in parameter space,
The Fisher metric is the Hessian of the KL divergence at zero separation.
Proof
Taylor-expand the log-likelihood around :
Substituting into the definition of KL divergence:
The first term vanishes because (the score has mean zero). For the second term, recall the Bartlett identity: . Therefore
The KL divergence is not a distance (it is asymmetric), but its symmetrised version has the same leading-order expansion . The Fisher metric captures the local, symmetric part of any divergence in the broad family of -divergences. This is why it is canonical: all reasonable notions of statistical distance agree infinitesimally, and they all agree with the Fisher metric.
4.5. Worked examples: the geometry of familiar models
The Bernoulli manifold
The Bernoulli family has Fisher metric
We show that this one-dimensional Riemannian manifold is isometric to a semicircle of unit radius. Define the Fisher-Rao coordinate , so that . Then
Substituting:
In the coordinate , the metric is flat: . As ranges over , ranges over . The total geodesic length is , and the geodesic distance between and is
Geometrically, parametrises a semicircular arc of radius by arc length: the embedding traces out a semicircle of unit radius in with total arc length .
The Bernoulli family equipped with the Fisher metric is isometric to the open semicircular arc of unit radius, via the map .
Proof
The computation in the example above shows that the map pulls back the flat metric to the Fisher metric . Since this map is a smooth bijection from to with smooth inverse , it is a diffeomorphism. A diffeomorphism that preserves the metric is an isometry. The interval with the flat metric is isometric to a semicircular arc of unit radius (parametrised by arc length), completing the proof.
The Normal manifold
The Normal family is a two-dimensional statistical manifold with coordinates . Its Fisher metric is
This is (up to a constant rescaling) the Poincare half-plane metric, the standard model of two-dimensional hyperbolic geometry.
We now compute the full Riemannian geometry of this surface: Christoffel symbols, geodesics, and curvature.
Christoffel symbols. The inverse metric is , , . Applying the Christoffel formula with coordinates , we note that all metric components depend only on (not on ), so any derivative . The nonzero derivatives are:
Computing each Christoffel symbol (all unlisted ones vanish):
from sympy import symbols, Matrix, simplify, Rational, diff
mu, sigma = symbols('mu sigma', positive=True)
# Fisher metric for N(mu, sigma): g = diag(1/sigma^2, 2/sigma^2)
g = Matrix([[1/sigma**2, 0], [0, 2/sigma**2]])
g_inv = g.inv()
coords = [mu, sigma]
print("Fisher metric g:")
print(g)
print("\nChristoffel symbols:")
for k in range(2):
for i in range(2):
for j in range(i, 2):
val = 0
for l in range(2):
val += Rational(1,2) * g_inv[k,l] * (
diff(g[j,l], coords[i]) +
diff(g[i,l], coords[j]) -
diff(g[i,j], coords[l])
)
val = simplify(val)
if val != 0:
print(f" Gamma^{k+1}_{{{i+1}{j+1}}} = {val}")Christoffel symbols for the Normal statistical manifold
Gaussian curvature. We compute using the definition of the Riemann tensor:
Wait: let us be precise with the index convention. Using , we want with :
(All other -terms vanish because the relevant Christoffel symbols are zero.) Therefore:
Lowering the index: . The determinant of the metric is . The Gaussian curvature is:
The Normal statistical manifold has constant negative Gaussian curvature . This is the hallmark of hyperbolic geometry: triangles have angle deficit, parallel geodesics diverge, and circles grow exponentially.
from sympy import symbols, Matrix, simplify, Rational, diff
mu, sigma = symbols('mu sigma', positive=True)
g = Matrix([[1/sigma**2, 0], [0, 2/sigma**2]])
g_inv = g.inv()
coords = [mu, sigma]
# Compute Gamma^k_ij
def christoffel(k, i, j):
val = 0
for l in range(2):
val += Rational(1,2) * g_inv[k,l] * (
diff(g[j,l], coords[i]) + diff(g[i,l], coords[j]) - diff(g[i,j], coords[l]))
return simplify(val)
# Riemann tensor R^1_212
R1_212 = (diff(christoffel(0,1,1), coords[0])
- diff(christoffel(0,0,1), coords[1]))
for m in range(2):
R1_212 += christoffel(0,0,m)*christoffel(m,1,1) - christoffel(0,1,m)*christoffel(m,0,1)
R1_212 = simplify(R1_212)
# R_1212 = g_1l R^l_212
R_1212 = simplify(g[0,0]*R1_212)
detg = simplify(g.det())
K = simplify(R_1212 / detg)
print(f"R^1_212 = {R1_212}")
print(f"R_1212 = {R_1212}")
print(f"det(g) = {detg}")
print(f"Gaussian curvature K = {K}")Gaussian curvature of the Normal statistical manifold
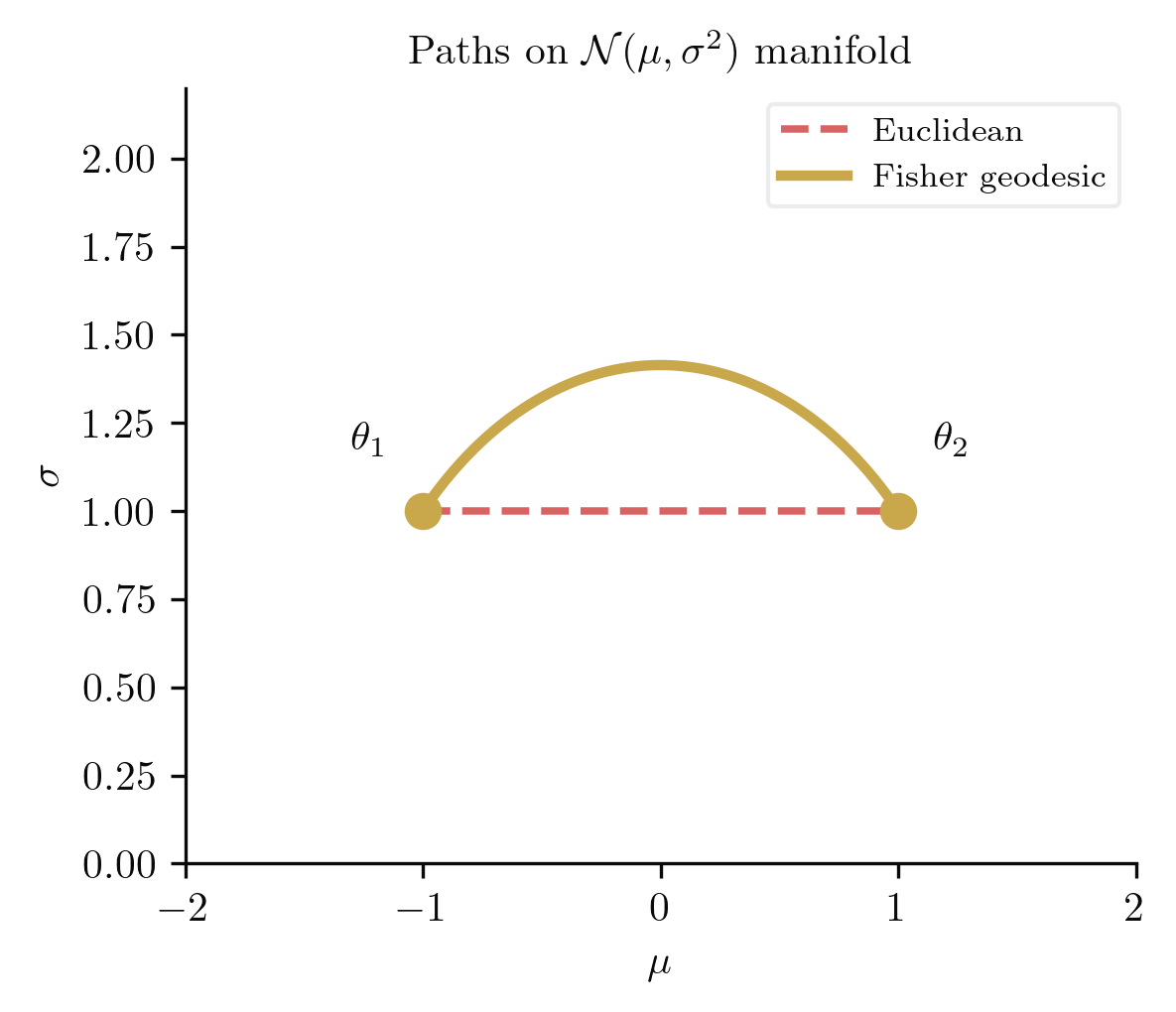
Geodesics. The geodesic equations for the Normal manifold are, writing and :
These are the geodesic equations of the Poincare half-plane (with the coordinate rescaling reducing to the standard form). The solutions are of two types: vertical lines (corresponding to pure changes in variance) and semicircles centred on the -axis ( boundary). In statistical terms, the geodesic between two Normal distributions curves upward through higher-variance regions, because at larger the distributions are more spread out and therefore closer together in Fisher distance.
4.6. The Bayesian-frequentist synthesis
The geometric viewpoint does more than provide elegant computations: it reveals that the Bayesian and frequentist frameworks are two coordinate systems on the same Riemannian manifold. Their convergence at large sample sizes is a geometric theorem.
Let for some interior point , and let be a prior that is continuous and positive at . Under standard regularity conditions, the posterior distribution satisfies
Geometrically: the posterior concentrates in a Fisher-metric ball of radius around the MLE. The shape of the ball is determined by the Fisher metric, not by the prior.
This theorem is the bridge. At large , Bayesian credible sets and frequentist confidence sets coincide, and both are determined by the Fisher geometry.
The Jeffreys prior is the prior distribution
where is the Fisher metric. It is the Riemannian volume form of the statistical manifold, and it is invariant under reparametrisation: if , then .
Jeffreys prior answers the question: what does "uniform" mean on a curved space? On a Riemannian manifold, the natural volume element is , not the coordinate volume . Integrating the constant function against the Riemannian volume gives the Riemannian volume of the region, not its coordinate volume. Jeffreys prior says: assign equal probability to equal volumes of statistical distance, not equal volumes of coordinate distance. In flat space the two coincide; on a curved statistical manifold they do not, and the Jeffreys prior automatically compensates for the curvature.
Bernoulli. The Fisher metric is , so and
This is the density of the distribution, also known as the arcsine distribution. It concentrates mass near and , precisely the regions where the Bernoulli manifold curves most sharply (where Fisher information is largest).
Normal. The Fisher metric is , so and
This is the standard reference prior for the Normal model. It is improper (does not integrate to a finite value), but it yields proper posteriors when combined with any non-degenerate likelihood.
The visualisation above shows the Bernstein-von Mises theorem in action. At small , the prior matters: the Bayesian credible interval (gold) and frequentist confidence interval (blue) diverge. At large , both converge to the same interval, centred at the MLE, with width determined by the Fisher information. The geometric viewpoint explains why: both frameworks respect the Fisher metric. The frequentist approach conditions on and averages over data; the Bayesian approach conditions on data and integrates over . But the Fisher metric governs the precision of both. Where they differ is in what they condition on, not in the geometry they inhabit.
4.7. Looking ahead
The Fisher metric is the beginning, not the end, of information geometry. The full theory, developed by Amari and others, introduces -connections: a one-parameter family of connections on the statistical manifold, of which the Levi-Civita connection is the member. The and connections (the e-connection and m-connection) are dual with respect to the Fisher metric, and exponential families are dually flat with respect to this pair. This dual structure governs not just distance but divergence, projection, and the geometry of model selection. That is the subject of the sequel.
References
- Kolmogorov, A. N. (1933). Grundbegriffe der Wahrscheinlichkeitsrechnung. Springer. English translation: Foundations of the Theory of Probability, 2nd ed., Chelsea, 1956. DOI
- Fisher, R. A. (1922). On the Mathematical Foundations of Theoretical Statistics. Philosophical Transactions of the Royal Society A, 222(594-604), 309-368. DOI
- Fisher, R. A. (1925). Theory of Statistical Estimation. Mathematical Proceedings of the Cambridge Philosophical Society, 22(5), 700-725. DOI
- Neyman, J. and Pearson, E. S. (1933). On the Problem of the Most Efficient Tests of Statistical Hypotheses. Philosophical Transactions of the Royal Society A, 231(694-706), 289-337. DOI
- Rao, C. R. (1945). Information and the Accuracy Attainable in the Estimation of Statistical Parameters. Bulletin of the Calcutta Mathematical Society, 37, 81-91. Link
- Cramer, H. (1946). Mathematical Methods of Statistics. Princeton University Press. Publisher
- Amari, S. and Nagaoka, H. (2000). Methods of Information Geometry. Translations of Mathematical Monographs, Vol. 191, AMS. DOI
- Amari, S. (2016). Information Geometry and Its Applications. Applied Mathematical Sciences, Vol. 194, Springer. DOI
- Lehmann, E. L. and Casella, G. (1998). Theory of Point Estimation, 2nd ed. Springer. DOI
- Lehmann, E. L. and Romano, J. P. (2005). Testing Statistical Hypotheses, 3rd ed. Springer. DOI
- van der Vaart, A. W. (1998). Asymptotic Statistics. Cambridge University Press. DOI
- Jeffreys, H. (1946). An Invariant Form for the Prior Probability in Estimation Problems. Proceedings of the Royal Society A, 186(1007), 453-461. DOI
- Le Cam, L. (1986). Asymptotic Methods in Statistical Decision Theory. Springer. DOI
- do Carmo, M. P. (1992). Riemannian Geometry. Birkhauser. DOI
- Efron, B. (1975). Defining the Curvature of a Statistical Problem (with Applications to Second Order Efficiency). Annals of Statistics, 3(6), 1189-1242. DOI
- Kullback, S. and Leibler, R. A. (1951). On Information and Sufficiency. Annals of Mathematical Statistics, 22(1), 79-86. DOI